Exploring the Medicinal Potential of Jatropha gossypifolia Leaf Extract: a Comprehensive Analysis of Bioactive Compounds, Antimicrobial Activities, and Molecular Interactions in the Treatment of Atopic Dermatitis
Abstract
Background:Jatropha gossypifolia, a plant traditionally recognized for its therapeutic properties, has gained scientific interest due to its potential pharmacological applications. However, comprehensive studies exploring its bioactive constituents and biomedical relevance remain limited. The aim of the study:To evaluate the phytochemical compositions, antimicrobial efficacy, and molecular interactions of Jatropha gossypifolia ethanolic leaf extract, with a particular focus on its potential role in managing atopic dermatitis. Materials and methods: The ethanolic extract of Jatropha gossypifolia leaves was analysed using GC-MS/MS to identify its bioactive compounds. Antimicrobial activity was assessed against clinical microbial isolates using standard in vitro methods. Molecular docking studies were performed in order to evaluate the binding affinities of key phytochemicals particularly Squalene and Phytol against target proteins involved in atopic dermatitis, including Interleukin-13, Interleukin-22, and JAK1 kinase. Results: GC-MS/MS analysis identified several bioactive compounds, notably Squalene and Phytol. The extract exhibited potent activity against diverse bacterial and fungal species. Molecular docking results revealed a strong binding affinity of Squalene with the selected target proteins, indicating its potential role in modulating inflammatory pathways associated with atopic dermatitis. Conclusion: This study highlights the therapeutic potential of Jatropha gossypifolia leaves, particularly its antimicrobial properties and its potential in targeting molecular pathways linked to atopic dermatitis. The findings support the necessity of further investigation into its development as a source of novel pharmacological agents for modern healthcare applications
Keywords: antimicrobial activities, atopic dermatitis, Jatropha gossypifolia, GC-MS/MS analysis, molecular docking, medicinal potential
Introduction The Euphorbiaceae family member Jatropha gossypifolia has garnered a lot of interest lately because of its varied phytochemical makeup and traditional medical applications. Commonly referred to as "Physic Nut" or "Bellyache Bush", this evergreen shrub has been used for generations in conventional medical practices in many different countries. The name is derived from the Greek words "jatros", meaning doctor, and "trophe", referring to nourishment. "Jatropha" has traditionally been utilized for medicinal purposes [1].
The fascination with the bioactive components of the plant prompted the necessity for a comprehensive phytochemical examination. Our focus revolves around identifying and quantifying secondary metabolites in Jatropha gossypifolia, employing cutting-edge methodologies [2]. This analysis forms the foundation for comprehending the plant's medicinal attributes and potential applications in modern medicine. To unravel the chemical composition, GC-MS/MS was employed, providing unparalleled insights into the diverse compounds that contribute to the plant's medicinal qualities. The relevance of its antimicrobial properties is accentuated against the backdrop of escalating antibiotic resistance [3]. Our study delves into the plant's efficacy against various microbial strains, exploring its potential as a source of antimicrobial drugs.
To clarify potential interactions between bioactive chemicals and important biomolecular targets, molecular docking experiments were carried out. This computational approach, estimating affinity and binding patterns, complements experimental findings and enhances our understanding of the plant's pharmacological actions [4]. The comprehensive synthesis of data from multiple approaches contributes to a nuanced comprehension of the medicinal potential of the plant.
Our study aims to identify synergistic relationships among the plant's constituents by correlating phytochemical profiles with biological activities. This holistic approach provides a comprehensive perspective on the medicinal capabilities of Jatropha gossypifolia leaves. The results highlight its potential to address contemporary healthcare challenges, inspiring optimism for the development of innovative medicinal agents. Specifically, the study delves into dermatological care, offering insights into the plant’s potential in treating atopic dermatitis [5, 6]. The integration of traditional knowledge with contemporary scientific investigation positions this research as a cornerstone for the development of new and natural therapies, aiming to provide comprehensive solutions for the intricate challenges associated with atopic dermatitis.
Materials and Methods
Sample collection
The fully grown, recently fresh, and untainted foliage of Jatropha gossypifolia was gathered from different locations in Eachanari, Coimbatore, Tamil Nadu. The leaves were thoroughly washed under running water and were subsequently air-dried in the shade for further utilization.
Preparation of Extract
The plant extract was produced using the solvent extraction method. First, the plant leaves were dried and ground into a powder using a mortar and pestle. The resulting powder was then sealed in filter paper and extracted in a Soxhlet extraction apparatus, each cycle using 200 ml of 100% ethanol as the solvent at 60° C [7, 8]. Further the extract underwent a rotary evaporation to eliminate undesired residues, yielding a concentrated extract. The resulting extract was condensed and stored 4°C for activity determination and analysis.
GC-MS/MS Analysis
The investigation of different bioactive substances within Jatropha gossypifolia leaf extract was accomplished through a comprehensive GC-MS/MS analysis. The evaluation utilized an Agilent make CH-GCMSMS02, 8890 GC System, and 7000 GC/featuring columns with dimensions of 30 m x 250µ x 0.25µ. Through an appropriate interface, the GC system was linked to the MS-D to facilitate mass spectrum detection. The temperature was planned to rise from its starting setting of 100°C to a final temperature of 270° C at a pace of 10°C per minute. The carrier gas utilized throughout the analysis was helium [9, 10] while the solvent system comprised Diluent-Ethanol. The entire analytical procedure for each sample spanned a total running time of 38 minutes.
Anti-bacterial activity
The antibacterial efficacy of Jatropha gossypifolia leaf extract was assessed using the disc diffusion method against clinical isolates of Streptococcus aureus, Salmonella typhi, and Pseudomonas aeruginosa obtained from CBNR, Eachanari, Coimbatore. The bacterial cultures were evenly spread on individual Muller Hinton agar plates and allowed to dry. Ampicillin (10µg) antibiotic disc, were employed for all three cultures [8]. Shortly before the experiment, the antibiotic discs were impregnated with plant extract at various concentrations (50 and 100 mg/mL) and placed on the agar surface. Ethanol was completely removed from the extract by evaporation and a solution of the extract was made by introducing 0.1g of extract with DMSO to constitute a final concentration of 100mg/mL. Then, a part of the stock was diluted to 50mg/mL concentration. 20 µL of the 50 and 100mg/mL were impregnated separately on the sterile discs, with 5 µL of extract being flecked on either sides of the discs and left to dry before the next set of 5 µL was spotted on the discs for optimal impregnation [9]. Along with two impregnated discs, each plate included a positive control (standard antibiotic disc) and a negative control (100% DMSO). Subsequently, the plates underwent incubation, and the resulting inhibition zones were examined.
Anti-fungal activity
The antifungal effectiveness was evaluated against a clinical isolate of Candida albicans collected from CBNR, Eachanari, Coimbatore. For the assay, fluconazole was employed as the standard antifungal agent. The fungal culture was evenly distributed on PDA agar plates. Along with the positive control, each plate featured a negative control (disc with 100% DMSO), and two previously impregnated discs, one immersed in the 50 mg/mL concentration of the stock solution and the other in the 100 mg/mL concentration, were used [10]. The plates were examined closely to check for any resulting inhibitory zones after incubation.
Docking studies
In our analysis, we conducted a docking study for compounds identified through GC-MS/MS analysis against proteins associated with atopic dermatitis. Specifically, the proteins involved in the disease are Interleukin-13, Interleukin-22, and JAK1 kinase. To acquire the 3D structures of the compounds, we utilized the PubChem database, while the 3D structures of the proteins were acquired from the Protein Data Bank. Protein preparation was done using Discovery Studio Visualizer 3.0, and the docking analysis was completed with PyRx, both of which are open-source tools for virtual screening.
Ligand Preparation
After conducting a screening based on Lipinski rule from the compounds screened through GC-MS/MS analysis [potential compounds were selected [11]. The 3D structures of phytol and squalene were obtained in the SDF format from the PubChem database.
Protein Preparation
Proteins associated with atopic dermatitis were selected based on existing literature. The 3D structures of IL-13 (PDB ID: 3LB6), IL-22 (PDB ID: 1YKB), and JAK1 kinase (PDB ID: 6N77) were obtained from the Protein Data Bank. The structures were prepared using Biovia software by removing any hydrogen atoms, heteroatoms or co-crystallised proteins present in the complex. Polar hydrogens were then added, and the final structures were saved in the PDB format following the protocol described by Mun et al. (2022). This preparation was necessary to reduce molecular complexity and ensure accurate docking, as crystallographic structures often lack hydrogen atom information. The same procedure was applied to all selected proteins prior to the docking analysis.
Docking
The docking study was performed using Pyrx software. For this process, the prepared protein was incorporated into Pyrx by using the open tool, designating the protein as the macromolecule. The compounds were introduced into Pyrx using Open Babel, and preparations for docking were made. Subsequently, energy minimization was performed for the ligands and protein, which will be effective during interaction. It was done as per the Universal Force Field, where it reduced energy to the minimum to make the bonds strong. The compounds were converted to the PDBQT format using the Open Babel tool. The docking procedure commenced with the use of Vina Wizard. Initially, we selected the protein, followed by the choice of compounds for screening. Further, the grid box was adjusted to the active sites of the proteins, where the ligands were expected to interact. The docking parameters were then adjusted to the default setting to ensure an effective docking process. The docking process was initiated [12-14]. After the docking was completed; the interactions of each compound were assessed by considering binding energy (kcal/mol). For every component, the binding pose with the lowest binding energy was chosen. To comprehend and visualize these interactions, Biovia Discovery Studio software were used for the analysis of the chosen compounds and their respective targets. This process was replicated for all three proteins, and the results were thoroughly discussed.
Result and discussion
GC-MS/MS Analysis
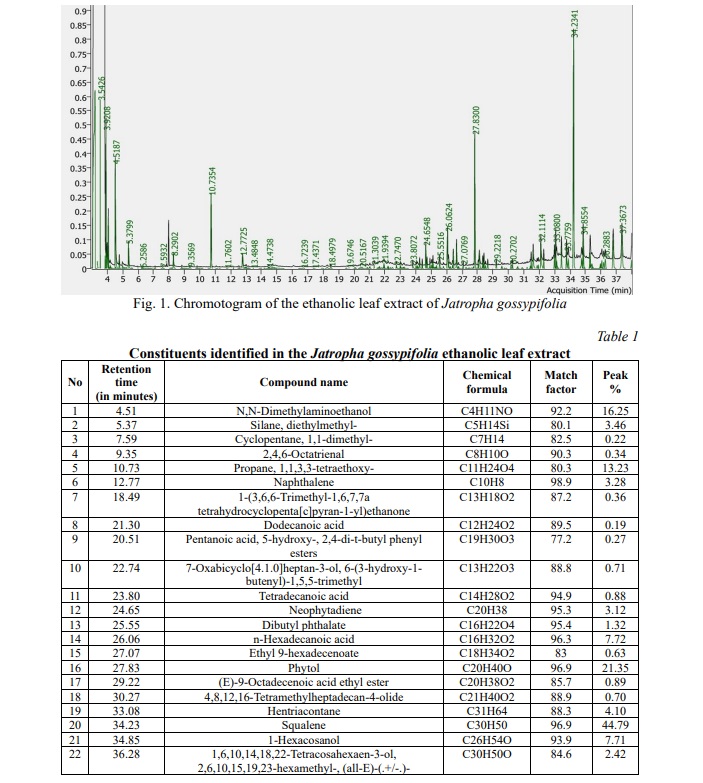
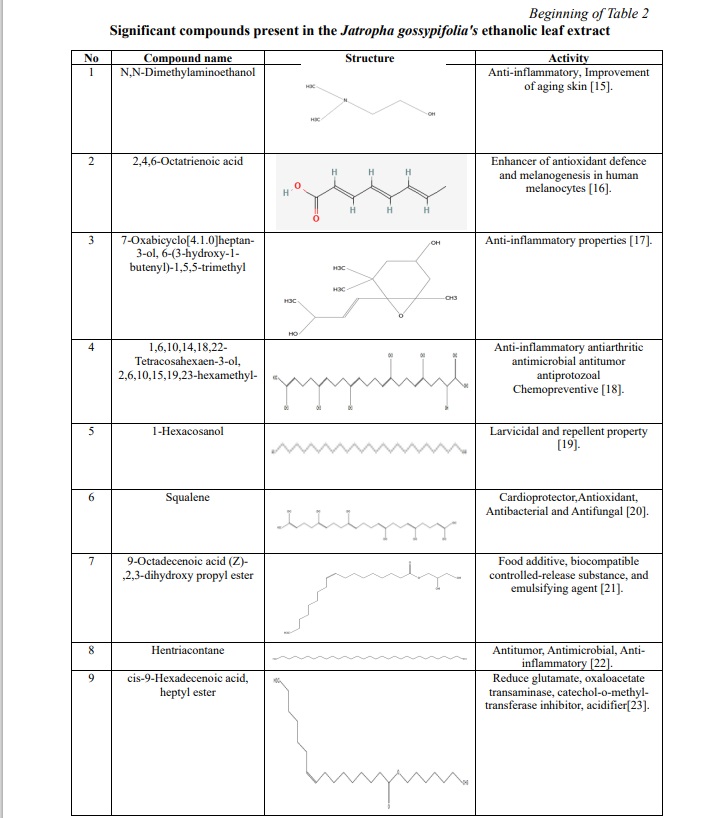
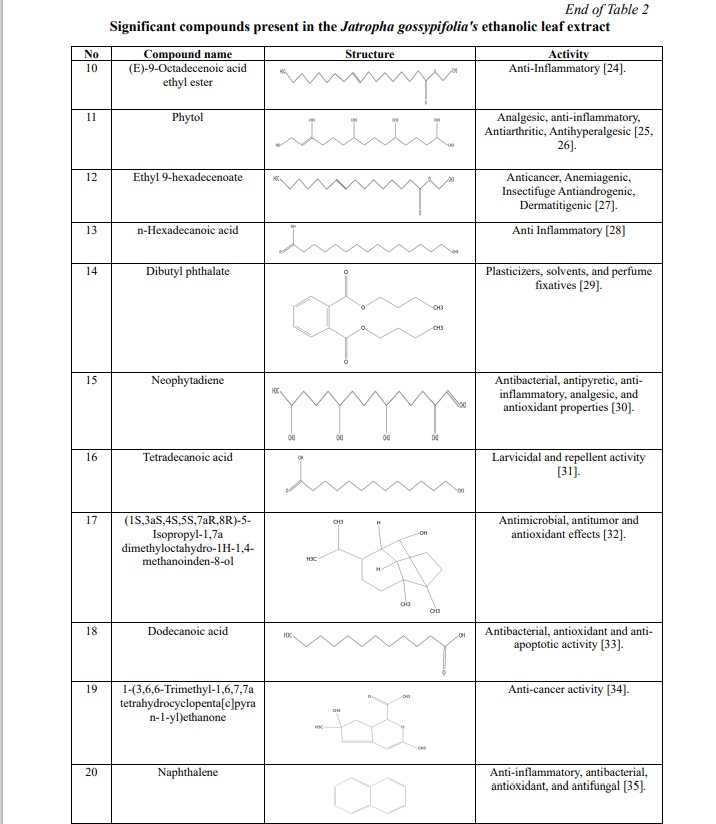
The results of the plant extract analysis revealed the existence of a number of compounds, which are displayed in Table 1, 2; Fig. 1. Furthermore, the compounds were identified based on their molecular weight and retention time, as compared to the NIST library. Among the list of compounds, squalene and phytol were the prominent compounds in our extract which has anti-inflammatory, antimicrobial, antibacterial properties.
Antibacterial activity
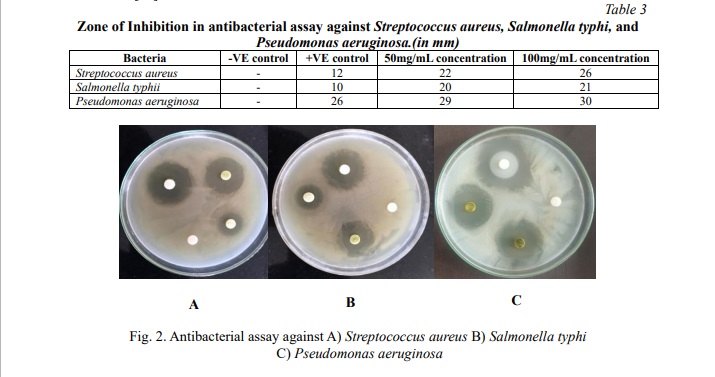
The ethanolic leaf extract of Jatropha gossypifolia exhibited significant activity [36] against pathogenic isolates of Streptococcus aureus, Salmonella typhi, and Pseudomonas aeruginosa, which were studied by the disc diffusion method. Microbial activities were quantitatively assessed based on the diameter of the inhibition zone (in millimeters) as shown in Fig. 2; Table 3. Jatropha gossypifolia shows the higher inhibition diameter for Pseudomonas aeruginosa of 30 mm for the 100 mg/mL concentration. Further, it shows the inhibition activity for all 3 species were similar in range and significant, which shows the efficiency of the antibacterial activity. Okoh et al. [37] demonstrated that the essential oil from Jatropha gossypifolia exhibits weaker bactericidal activity against Gram-negative bacteria than Gram-positive ones. Conversely, this study found that the ethanolic extract exhibited significant bactericidal activity against both Gram-positive and Gram-negative bacteria. This result can be due to the specific chemical composition and driving forces. Many essential oils have a high content of lipophilic terpenes and aromatics and therefore may have relatively little capacity to penetrate the outer membrane of Gram-negative bacteria. In contrast, ethanolic extracts have a wider range of polar components, and can penetrate easily through the dual membrane of Gram-negative bacteria, strengthening their antibacterial action.
Antifungal activity
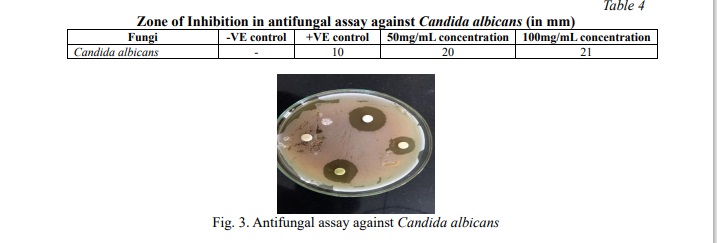
The ethanolic leaf extract of Jatropha gossypifolia leaves exhibited significant antifungal ability against the clinical isolate of Candida albicans. The inhibition zone was observed as shown in Table 4; Fig. 3. Antifungal activity of Jatropha gossypifolia shows inhibition with diameter of 21 mm for 100mg/ml concentration, which has higher than the positive control. Also, this assay shows that at 50mg/ml concentration, it has efficient inhibition of 20 mm diameter as like the 100 mg/ml concentration. In an earlier investigation, Félix-Silva et al. [38] observed only mild antifungal effects from the chloroform and hexane fractions of Jatropha gossypifolia. In contrast, this study shows that the ethanolic extract shows considerably stronger activity against Candida albicans. This difference may be due to the polarity of ethanol, enabling it to extract a wider range of active phytochemicals. Compounds such as squalene and phytol, present in our ethanolic extract, are likely contributors to this increased activity. These constituents are believed to interfere with the fungal cell membranes by disrupting lipid organization and inhibiting ergosterol biosynthesis. Based on these findings, ethanol appears to be a more effective solvent for isolating antifungal agents from J. gossypifolia than non-polar alternatives. This underscores the importance of solvent choice in phytochemical extraction and its impact on biological activity.
Docking studies
A docking study shows how ligands interact with proteins. The Pyrx tool can screen ligands with nine different poses of the target protein's active site to identify better interactions. The interactions of each compound were assessed based on binding energy (kcal/mol), and for every compound, the binding pose with the lowest energy was chosen. These energies depict the interaction of the compounds with proteins, which involved the protein mechanism to inhibit the inflammatory proteins. To gain insights into these interactions, the selected compounds and their targets were analyzed using Biovia Studio software, facilitating visualization and comprehension of the molecular interactions. This can visualize the interaction bonds and forces of the compounds with amino acid. Further, it represents the biological changes of the proteins during interaction. After docking, we acquired the results for three different proteins against 2 compounds which were obtained from GC-MS/MS analysis. (Table 5) represents the docking result for the Interleukin-13 (3LB6) protein against squalene and phytol. From this result, it is evident that squalene has the least binding affinity of -6.6 kcal/mol, indicating a more potent and robust interaction with the protein. (Fig. 4, 5) depict the 2D diagram illustrating the interaction between amino acids and the compound. The phytol compound interacted with Arg108 via a hydrogen bond, while Phe107, Arg11, Ile14 and Val18 interacted with the ligand via an alkyl bond. Lys and Glu interacted via van der Waals forces. Similarly, squalene strongly interacted with amino acids through strong alkyl bonds.
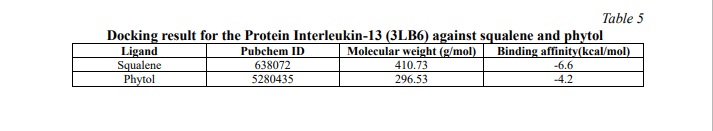
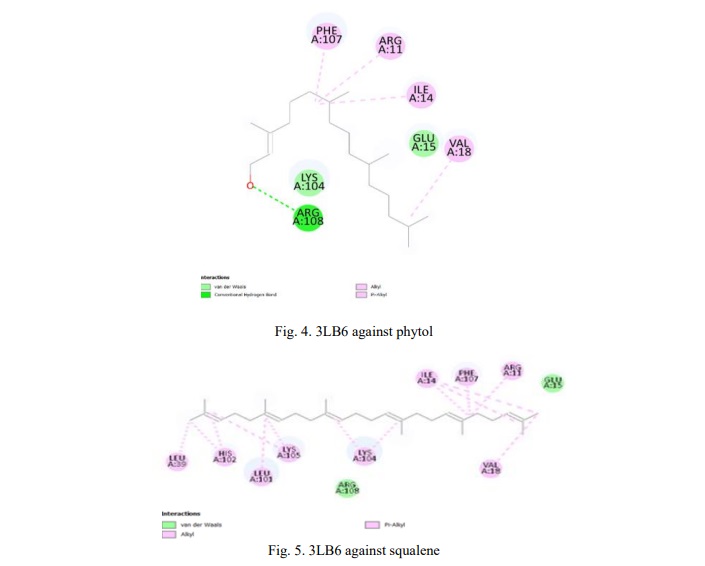
Table 6 shows the binding energy for the Interleukin-22 (1YKB) protein against the squalene and phytol compound. Based on this result, we concluded that both compounds exhibit a significant binding affinity of -5.1 kcal/mol. (Figures 6, 7) show the 2D diagram of the interaction between amino acids and compounds. Further investigation of the bonds and interaction forces between different amino acids and ligands was carried out. Squalene strongly interacted with the amino acid's hydrogen bonds, showing significant binding energy, while phytol interacted with the amino acid through van der Waals interactions.
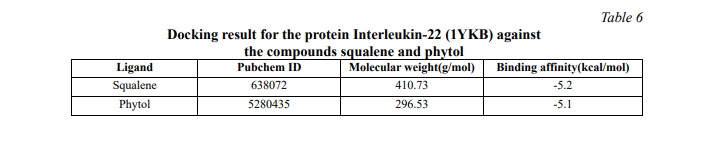
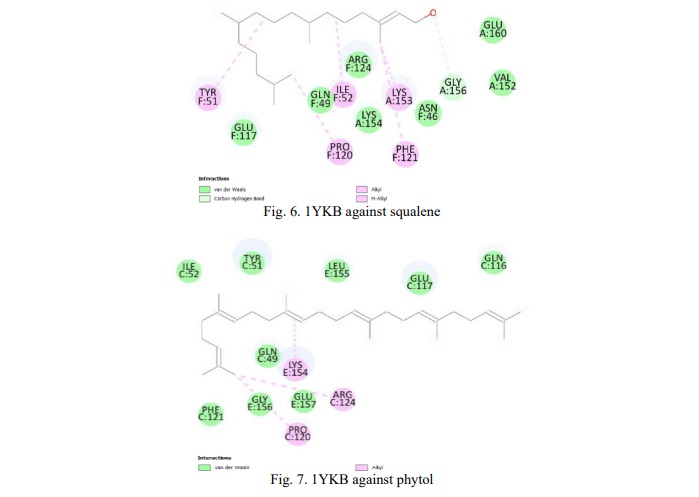
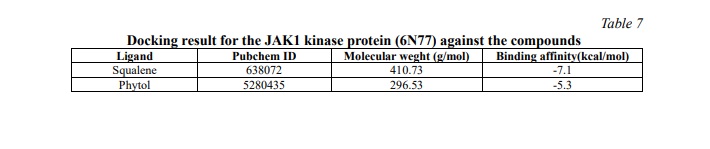
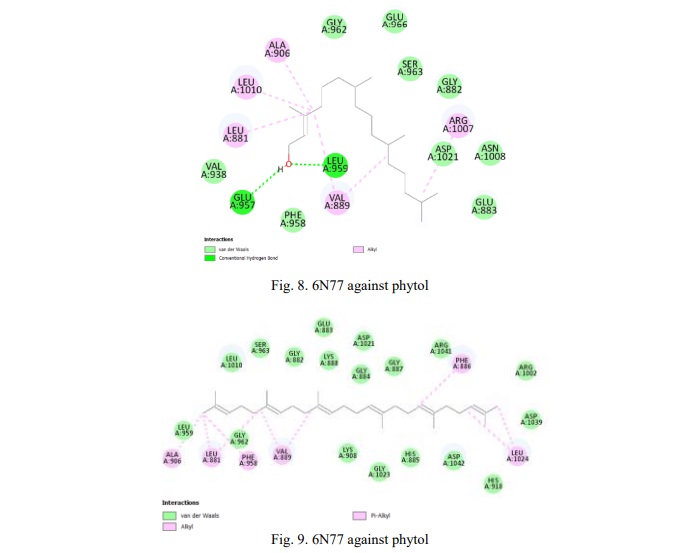
Table 7 shows the result for the JAK1 kinase protein against the compounds. The compound squalene has the least binding affinity of -7.1 kcal/mol which has more potential in targeting this protein. Fig. 8, 9 show the 2D representation of the interaction of 6N77 protein amino acid with the compounds. The figures illustrate the bonds and forces between the amino acid and ligands. Phytol interacts with amino acids through van der Waals forces and forms hydrogen bonds with the amino acid chains. Similarly, squalene was bound to the amino acid via alkyl bonds.
Conclusion. In conclusion, Jatropha gossypifolia, renowned for its rich phytochemical composition and historical medicinal uses, has been subject to a comprehensive study. Employing advanced techniques such as GC-MS/MS and molecular docking we uncovered the plant's diverse bioactive compounds and explored their potential in medicinal applications. Notably, our research highlights the antibacterial and antimicrobial properties of Jatropha gossypifolia, crucial in the face of increasing antibiotic resistance. The study's integrative approach, which combines traditional knowledge with modern scientific investigation, establishes Jatropha gossypifolia leaves as a promising source of novel medicinal agents. Specifically, our focus on its potential to treat atopic dermatitis adds depth to research into dermatological care, offering prospects for innovative, natural therapies. In vivo and in vitro studies are essential for improving our understanding of atopic dermatitis. Combining these approaches enhances our understanding of the complex pathophysiology and aids the development of effective treatments. Overall, this comprehensive study provides valuable insights into the multifaceted medicinal properties of Jatropha gossypifolia, paving the way for further exploration and potential advancements in healthcare.
Financial support
No financial support has been provided for this work.
Thanks
The authors gratefully acknowledge Dr. R. Ramachandra Ragunathan, Department of Biotechnology (Bio-nanotechnology), Centre for Bioscience and Nanoscience Research, for generously providing access to testing facilities. His support has been instrumental to the success of our study





















Reference lists